Config files explained¶
The aim of these config files is to automate running the weak lensing measurements and post-processing.
The module xpipe.paths automatically tries to read the configs upon import, but you can also specify them later.
In the pipeline mode xpipe keeps track of reading lens catalogs, splitting them up into parameter bins, measuring the lensing signal and estimating covariances and contaminations automatically. However in order to do this, first we need to specify the details of these tasks in the below config files:
params.yml defines the measurement parameters
inputs.yml defines the available data files, and short aliases for them
These files do not exist yet when you first clone the repository, however there is a
default_params.yml and default_inputs.yml which you should use as a reference. These defaults are set up such
that when you create params.yml and inputs.yml they will be automatically looked for and read.
Config files are defined as yaml files, and are read as dictionaries, each entry consisting of a key and a corresponding value (note that this includes nested dictonaries).
Note that in yaml one should use null instead of None
Load order¶
default_params.yml
looks for params.yml
tries to read
custom_params_filefrom params.yml
from this point the load is recursive, e.g. param files are loaded as long as there is a valid custom params file defined in the last loaded config. Each new config file only updates the settings, such that keys which are not present in the later files are left at their latest value.
The parameters defined here are loaded into the dictionary xpipe.paths.params
params.yml¶
Key reference
custom_params_file: params.ymlIf you want to use an other parameter file, then specify it here. It must be in the same directory
custom_data_path: FalseAbsolute path to the
datadirectory of the pipeline. If False: usesdefault project_path + /datamode: fullThe pipeline supports two modes:
fullanddev. This is primarily used in setting up the input files for the measurement. e.g. you can define two binning schemes: one really complex for the full run, and a simple, quicker for devtag: defaultPrefix for all files (with NO trailing “_”). In addition this will be the name of the directory wher input and output files are written to.
shear_style: reducedFormat of the source galaxy catalog. Available formats are
reduced,lensfitandmetacalcat_to_use: defaultAlias for the lens catalog to be used (in this case the
default). Aliases are defined in inputs.ymlshear_to_use: defaultAlias for the source catalog to be used (in this case the
default). Aliases are defined in inputs.ymlparam_bins_fullParameter bins defined for
mode: full, e.g.:param_bins_full: q0_edges: [0.2, 0.35, 0.5, 0.65] q1_edges: [5., 10., 14., 20., 30., 45., 60., 999]
q0andq1refer to the zero-th and first quantities (in this order) you want to split your lens catalog by. For defining what these relate to seelenskeyandrandkey. In the above exampleq0is redshift, andq1is optical richness.In general you can define an arbitrary number of quantities keeping the notation that the binning edges for quantity n are written as
q[n]_edges.param_bins_devParameter bins defined for
mode: dev, e.g.:param_bins_dev: q0_edges: [0.2, 0.35] q1_edges: [45, 60]
q0andq1refer to the zero-th and first quantities (in this order) you want to split your lens catalog by. For defining what these relate to seelenskeyandrandkey. In the above exampleq0is redshift, andq1is optical richness.In general you can define an arbitrary number of quantities keeping the notation that the binning edges for quantity n are written as
q[n]_edges.lenskeyAliases for the columns of the lens data table (assuming fits-like record table):
lenskey: id: MEM_MATCH_ID ra: RA dec: DEC z: Z_LAMBDA q0: Z_LAMBDA q1: LAMBDA_CHISQ
q0andq1refer to the zero-th and first quantities (in this order) you want to split your lens catalog by (seeparam_bins_*). In general you can define an arbitrary number of quantities keeping the notation that the alias for quantity n are written asq[n]. In the above exampleq0is redshift, andq1is optical richness.randkeyAliases for the columns of the random points data table (assuming fits-like record table):
randkey: q0: ZTRUE q1: AVG_LAMBDAOUT ra: RA dec: DEC z: ZTRUE w: WEIGHT
q0andq1refer to the zero-th and first quantities (in this order) you want to split your random points catalog by. In general you can define an arbitrary number of quantities keeping the notation that the alias for quantity n are written asq[n]. In the above exampleq0is redshift, andq1is optical richnessNote that for random points you have to specify the same quantities as for the lens catalog.
nprocess: 2Number of maximum processes or CPU-s to use at the same time (OpenMP-style parallelization).
njk_max: 100Maximum number of Jackknife regions to use in resampling. Actual number is
max(n_lens, njk_max)nrandomsNumber of random points to use:
nrandoms: full: 50000 dev: 1000
seedsRandom seed for choosing the random points
random_seed, and for generating rotated shear catalogsshear_seed_master:seeds: random_seed: 5 shear_seed_master: 10
cosmo_paramsCosmology parameters defined as:
cosmo_params: H0: 70. Om0: 0.3
radial_binsLogarithmic (base 10) radial bins from rmin to rmax:
radial_bins: nbin: 15 rmin: 0.0323 rmax: 30.0 units: Mpc
Available units:
Mpc,comoving_mpcorarcminweight_style: "optimal"Source weight style in the xshear lensing measurement. Use
optimalwhen estimating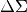 and
and uniformwhen measuring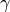 .
.pairlogSpecifies the amount of source-lens pairs to be saved, and for which radial range:
pairlog: pairlog_rmin: 0 pairlog_rmax: 0 pairlog_nmax: 0
Note that the pair limit is considered for each call of xshear separately. That is if you separate lenses into Jackknife regions then this is applicable for a single region.
lens_prefix: y1clustPrefix for lens-files
rand_prefix: y1randPrefix for random points files
subtr_prefix: y1subtrPrefix for lens - random points files
fields_to_use: ['spt', 's82']List of names of observational fields to use (as defined below)
fieldsDefinition of observational field boundaries:
fields: spt: dec_top: -30. dec_bottom: -60. ra_left: 0. ra_right: 360. s82: dec_top: 10. dec_bottom: -10. ra_left: 300. ra_right: 10. d04: dec_top: 10. dec_bottom: -30. ra_left: 10. ra_right: 250.
These can be approximate, the only requirement is that they divide the lens dataset into the appropriate chunks
pzparsParameters for the boost factor extraction:
pzpars: hist: nbin: 15 zmin: 0.0 zmax: 3.0 tag: "zhist" full: tag: "zpdf" boost: rbmin: 3 rbmax: 13 refbin: 14
There are two modes histogram
histwhich relies on Monte-Carlo samples of redshifts and is less robust, andfullwhich uses the full P(z) of each source galaxy.tagdefines the name appended to the corresponding files.boostdefines the radial range for the boost estimation in radial bins
pdf_paths: nullRegular expression matching the absolute paths of the
BPZoutput files containing the full redshift PDF. (e.g./home/data/*.h5).NOTE This is only required for estimating the Boost factors, and can be safely left
nullin a simple lensing run.
inputs.yml¶
This config file lists the available data products. Currently all products are listed under the local
key, indicating that they are found on disk, (as opposed to downloaded from some network location).
The two major sub-headings are:
shearcatLists the available xshear-style source catalog files located within:
[custom_data_path]/shearcat/
where
[custom_data_path]is the absolute path to thedatafolder specified by the corresponding key in params.ymlEach input file has it’s key as an alias for the file name, such that you can use the key you define here for a valid value of
shear_to_usefor params.yml, e.g.:shearcat: default: default.dat im3shape: im3shape_shear_catalog.dat metacal: metacal_shear_catalog.dat
These input files should be written in ASCII
lenscatLists the available lens catalog files located within:
[custom_data_path]/lenscat/
where
[custom_data_path]is the absolute path to thedatafolder specified by the corresponding key in params.ymlEach dataset has it’s key as an alias, which you can use to define the lens dataset for a valid value of
cat_to_usefor params.yml. In addition, each dataset is implicitely assumed to consist of a lens catalog, and a corresponding catalog of random points, such that for each key there are two sub-keys:lensandrand. Both of these files should be written in fits format:lenscat: y1clust: lens: des_y1_lens_catalog.fits rand: des_y1_rand_catalog.fits svclust: lens: des_sv_lens_catalog.fits rand: des_sv_rand_catalog.fits testclust: lens: test_catalog.fits rand: null
In case there are no random points available for the dataset you are using, it is safe to leave the
randfield empty, but in this case make sure you also use the--norandsflag when exectuing the pipeline scripts.In case the input catalog is defined in multiple files (for example when the parameter bins are not trivial to define), a list of filenames can be defined for
lensandrand:lenscat: pre_binned_data: lens: [ [des_y1_lens_catalog_bin-0-0.fits, des_y1_lens_catalog_bin-0-1.fits], [des_y1_lens_catalog_bin-1-0.fits, des_y1_lens_catalog_bin-1-1.fits], ]
Note: The defined files will be assumed to correspond to separat parameter selections, and thus this mode cannot be used together with the definiton of parameter bins in params.yml