xpipe.tools.selector.selector¶
- xpipe.tools.selector.selector(pps, limits)[source]¶
Applies selection to array based on the passed parameter limits
Selection is defined like
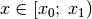
Examples
First make some mock data:
np.random.seed(seed = 5) nrows = 100000 ids = np.arange(nrows, dtype=int) ra = np.random.uniform(low=0., high=360., size=nrows) dec = np.random.uniform(low=-60., high=10., size=nrows) z = np.random.uniform(low=0.1, high=1.0, size=nrows) lamb = np.random.uniform(low=20, high=40, size=nrows) data = np.vstack((ra, dec, z, lamb)).T
then define binning edges:
ra_edges = (0., 100., 300.) dec_edges = (-60., -30., 10.) z_edges = (0.2, 0.4, 0.8) lamb_edges = (20., 30., 45., 60.) edges = (ra_edges, dec_edges, z_edges, lamb_edges)
This selection divides the data into 24 bins in 4-D space, which we can obtain perform as:
sinds, bounds, plpairs = sl.selector(data, edges)
Let’s inspect the output,
sindscontains the boolean indexing arrays to select rows by, whileboundsshows the corresponding parameter boundaries, for example:>>> bounds[13] ((0, (100.0, 300.0), 1), (1, (-60.0, -30.0), 0), (2, (0.2, 0.4), 0), (3, (30.0, 45.0), 1))
where the first number indicates the column of the data file it corresponds to, and the tuple contains the parameter boundaries for that column. The last element is an index for the original parameter edge. Indeed we find:
>>> data[sinds[13], :].min(axis=0) array([ 100.03881951, -59.96803764, 0.20010631, 30.02361767]) >>> data[sinds[13], :].max(axis=0) array([ 299.68649402, -30.00117535, 0.39994843, 44.99060231])
which are the boundaries we were expecting
- Parameters
pps (np.array N-D) – numpy array with N rows, containing the parameters to split the sample by
limits (list of lists) – list of parameter limits, each element contains limits for a columna of pps
- Returns
list of boolean indices (one for each selection), list of param limits corresponding to each selection parameter limits expanded from sequence to list of tuples
- Return type